---
title: "Correlation ABI, NPP and EVI"
format:
html:
toc: false
execute:
message: false
warning: false
editor_options:
chunk_output_type: console
---
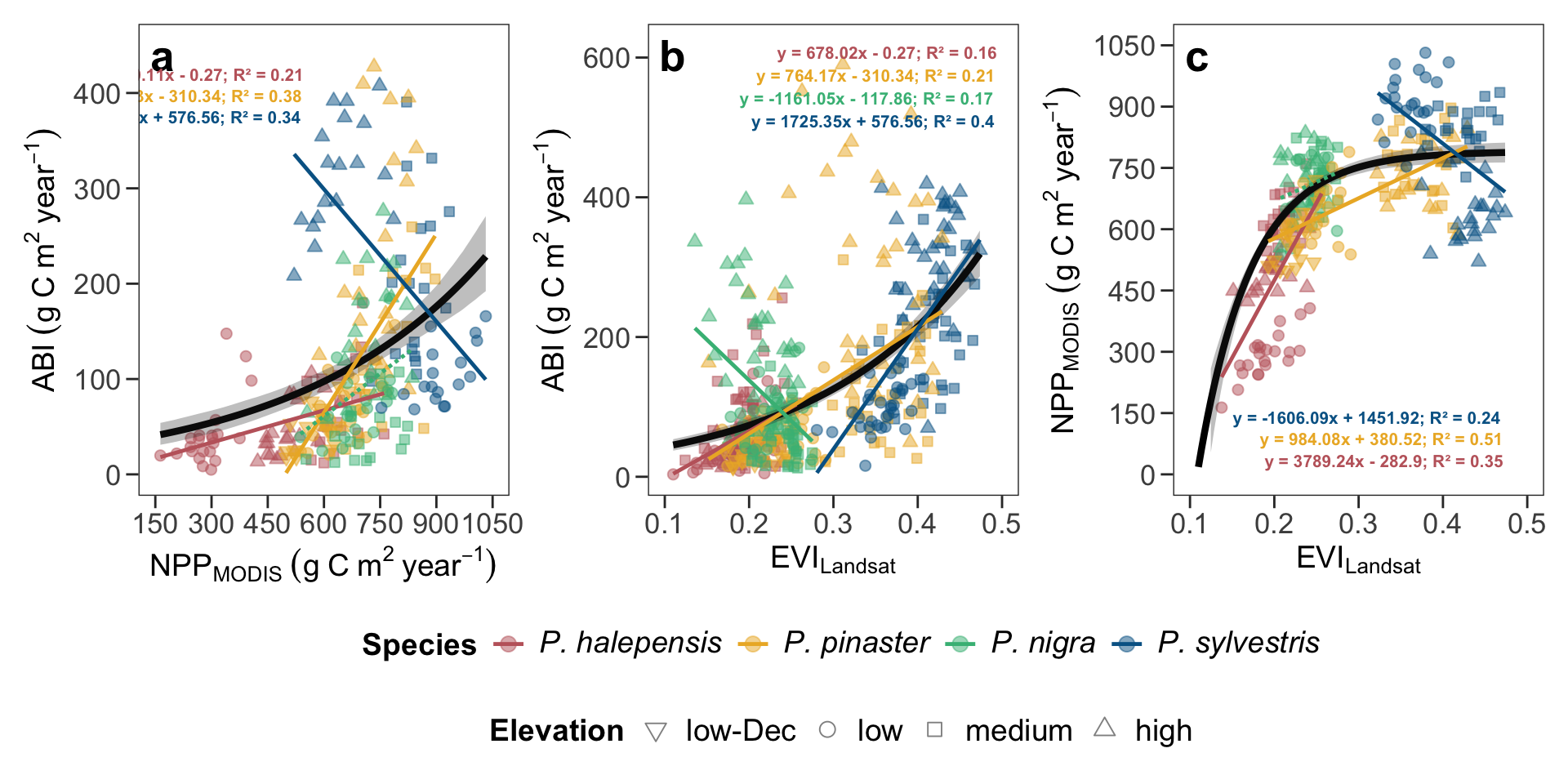
- This figure corresponds to figure 4 in the manuscript.
- An high-resolution version of this figure is available [here](../output/plot_compare_abi_npp_evi_sp.png).
```{r}
#| code-fold: true
#| fig-cap: "Relationships between Aboveground Biomass Increment (ABI) with: (a) annual Net Primary Productivity (NPP) (2001-2022); (b) annual Enhanced Vegetation Index (EVI) (1986-2022); and (c) between annual Net Primary Productivity (NPP) and annual Enhanced Vegetation Index (EVI) (2001-2022). The global trends for all pine species are depicted by a black line (gray shaded indicated bootstrap confidence intervals), while color-coded lines indicate relations for individual pine species (solid line: significant (p<0.001), dashed line: not significant)."
#| fig-width: 10
# pkgs
library(tidyverse)
library(boot)
library(patchwork)
library(here)
library(glue)
library(broom)
source(here::here("scripts/aux.R"))
options(scipen = 999)
# Read data
## A
abi <- read_csv(here::here("data/abi.csv")) |>
mutate(se = NA, sd = NA, variable = "abi") |>
rename(mean = abi)
## EVI Landsat
evi_landsat <- read_csv(here::here("data/iv_landsat.csv")) |>
filter(iv == "evi") |>
dplyr::select(year, sp_code, elev_code, sp_elev, mean, sd, se) |>
mutate(variable = "evi_landsat")
npp <- read_csv(here::here("data/npp_modis.csv")) |>
rename(mean = npp) |>
mutate(se = NA, sd = NA, variable = "npp") |>
dplyr::select(year, sp_code, elev_code, sp_elev, mean, sd, se, variable)
df_index <- bind_rows(
abi, evi_landsat, npp
) |>
mutate(elev_code = fct_relevel(elev_code, "low-Dec", "low", "medium", "high")) |>
mutate(sp_code = fct_relevel(sp_code, "halepensis","pinaster", "nigra", "sylvestris")) |>
mutate(Specie = paste0("P. ", sp_code)) |>
mutate(Specie = fct_relevel(Specie, "P. halepensis","P. pinaster", "P. nigra", "P. sylvestris")) |>
rename(mean_y = mean, y_variable = variable)
abi_npp <- df_index |>
filter(y_variable %in% c("abi", "npp")) |>
dplyr::select(-se, -sd) |>
pivot_wider(values_from = mean_y, names_from = y_variable) |>
filter(!is.na(npp)) |>
filter(!is.na(abi)) |>
rowwise() |>
mutate(ratio = abi / npp)
## ABI vs NPP
# Generate a confidence interval with bootstrap
# Define a function to perform the bootstrapping
boot_func_abi_npp <- function(data, indices) {
# Take a bootstrap sample
sample_df <- data[indices, ]
# Fit the non-linear model
nls_fit <- nls(abi ~ a * exp(b * npp), data = sample_df, start = list(a = 10, b = 0.002958839))
# Generate predicted values for a sequence of npp
npp_seq <- seq(min(data$npp), max(data$npp), length.out = 100)
pred <- predict(nls_fit, newdata = data.frame(npp = npp_seq))
return(pred)
}
# Perform bootstrapping with 1000 resamples
boot_res_abi_npp <- boot(data = abi_npp, statistic = boot_func_abi_npp, R = 1000) # R is the number of resamples
# Compute the 95% confidence intervals for each point
ci_lower <- apply(boot_res_abi_npp$t, 2, quantile, probs = 0.025) # Lower 2.5%
ci_upper <- apply(boot_res_abi_npp$t, 2, quantile, probs = 0.975) # Upper 97.5%
# Predicted values from the original fit
npp_seq <- seq(min(abi_npp$npp), max(abi_npp$npp), length.out = 100)
pred_abi_npp <- apply(boot_res_abi_npp$t, 2, mean) # Mean of the bootstrapped predictions
# Create a data frame with predicted values and confidence intervals
pred_df_abi_npp <- data.frame(
npp = npp_seq,
abi = pred_abi_npp,
ci_lower = ci_lower,
ci_upper = ci_upper
)
labi_npp <- abi_npp |>
group_by(sp_code) |>
group_modify(~ {
# Ajusta el modelo
mod <- lm(abi ~ npp, data = .x)
# Tidy con coeficientes
tidy_df <- broom::tidy(mod)
# Añade R2 a cada fila
tidy_df$r.squared <- broom::glance(mod)$r.squared
tidy_df
}) |>
mutate(p = case_when(
p.value < 0.001 ~ "<0.001",
p.value >= 0.001 ~ as.character(round(p.value, 3))
)) |>
mutate(sig = case_when(
p.value < 0.001 ~ "sig",
p.value >= 0.001 ~ "no sig"
))
y_max_abi <- 450
labi_npp_eq <- labi_npp |>
left_join(
labi_npp |> filter(term == "(Intercept)") |>
dplyr::select(sp_code, intercept = estimate)
) |>
mutate(
eq_text = if_else(
sig == "sig",
glue("y = {round(estimate, 2)}x {ifelse(intercept < 0, '-', '+')} {round(abs(intercept), 2)}; R² = {round(r.squared, 2)}"),
NA
)
) |>
filter(term != "(Intercept)") |>
arrange(desc(sig)) |> # ordenar por eq
rowid_to_column("id") |>
mutate(offset_y = id * 0.05 * y_max_abi,
y_max = y_max_abi - offset_y) |>
mutate(Specie = paste0("P. ", sp_code))
abi_npp <-
abi_npp |> inner_join(
labi_npp_eq|> dplyr::select(sp_code, p, sig))
## ABI vs EVI
abi_evi <- df_index |>
filter(y_variable %in% c("abi", "evi_landsat")) |>
dplyr::select(-se, -sd) |>
pivot_wider(values_from = mean_y, names_from = y_variable) |>
filter(!is.na(evi_landsat)) |>
filter(!is.na(abi))
# Generate a confidence interval with bootstrap
# Define a function to perform the bootstrapping
boot_func_abi_evi <- function(data, indices) {
# Take a bootstrap sample
sample_df <- data[indices, ]
# Fit the non-linear model
nls_fit <- nls(abi ~ a * exp(b * evi_landsat), data = sample_df, start = list(a = 15, b = 6.128))
# Generate predicted values for a sequence of npp
evi_seq <- seq(min(data$evi_landsat), max(data$evi_landsat), length.out = 100)
pred <- predict(nls_fit, newdata = data.frame(evi_landsat = evi_seq))
return(pred)
}
# Perform bootstrapping with 1000 resamples
boot_res_abi_evi <- boot(data = abi_evi, statistic = boot_func_abi_evi, R = 1000) # R is the number of resamples
# Compute the 95% confidence intervals for each point
ci_lower <- apply(boot_res_abi_evi$t, 2, quantile, probs = 0.025) # Lower 2.5%
ci_upper <- apply(boot_res_abi_evi$t, 2, quantile, probs = 0.975) # Upper 97.5%
# Predicted values from the original fit
evi_seq <- seq(min(abi_evi$evi_landsat), max(abi_evi$evi_landsat), length.out = 100)
pred_abi_evi <- apply(boot_res_abi_evi$t, 2, mean) # Mean of the bootstrapped predictions
# Create a data frame with predicted values and confidence intervals
pred_df_abi_evi <- data.frame(
evi_landsat = evi_seq,
abi = pred_abi_evi,
ci_lower = ci_lower,
ci_upper = ci_upper
)
# labi_evi <- abi_evi |>
# group_by(sp_code) |>
# group_modify(
# ~ broom::tidy(lm(abi ~ evi_landsat, data = .x))
# ) |>
# filter(term != "(Intercept)") |>
# mutate(p = case_when(
# p.value < 0.001 ~ "<0.001",
# p.value >= 0.001 ~ as.character(round(p.value, 3))
# )) |>
# mutate(sig = case_when(
# p.value < 0.001 ~ "sig",
# p.value >= 0.001 ~ "no sig"
# ))
labi_evi <- abi_evi |>
group_by(sp_code) |>
group_modify(~ {
# Ajusta el modelo
mod <- lm(abi ~ evi_landsat, data = .x)
# Tidy con coeficientes
tidy_df <- broom::tidy(mod)
# Añade R2 a cada fila
tidy_df$r.squared <- broom::glance(mod)$r.squared
tidy_df
}) |>
mutate(p = case_when(
p.value < 0.001 ~ "<0.001",
p.value >= 0.001 ~ as.character(round(p.value, 3))
)) |>
mutate(sig = case_when(
p.value < 0.001 ~ "sig",
p.value >= 0.001 ~ "no sig"
))
y_max_abi <- 650
labi_evi_eq <- labi_evi |>
left_join(
labi_npp |> filter(term == "(Intercept)") |>
dplyr::select(sp_code, intercept = estimate)
) |>
mutate(
eq_text = if_else(
sig == "sig",
glue("y = {round(estimate, 2)}x {ifelse(intercept < 0, '-', '+')} {round(abs(intercept), 2)}; R² = {round(r.squared, 2)}"),
NA
)
) |>
filter(term != "(Intercept)") |>
arrange(desc(sig)) |> # ordenar por eq
rowid_to_column("id") |>
mutate(offset_y = id * 0.05 * y_max_abi,
y_max = y_max_abi - offset_y) |>
mutate(Specie = paste0("P. ", sp_code))
abi_evi <-
abi_evi |> inner_join(
labi_evi_eq |> dplyr::select(sp_code, p, sig))
## NPP vs EVI
npp_evi <- df_index |>
filter(y_variable %in% c("npp", "evi_landsat")) |>
dplyr::select(-se, -sd) |>
pivot_wider(values_from = mean_y, names_from = y_variable) |>
filter(!is.na(evi_landsat)) |>
filter(!is.na(npp))
boot_func_npp_evi <- function(data, indices) {
# Take a bootstrap sample
sample_df <- data[indices, ]
# Fit the non-linear model
nls_fit <- nls(npp ~ Asym + (R0 - Asym) * exp(-exp(lrc) * evi_landsat),
data = sample_df,
start = list(Asym = 700, R0 = -6000, lrc = 2.8)
)
# Generate predicted values for a sequence of npp
evi_seq <- seq(min(data$evi_landsat), max(data$evi_landsat), length.out = 100)
pred <- predict(nls_fit, newdata = data.frame(evi_landsat = evi_seq))
return(pred)
}
# Perform bootstrapping with 1000 resamples
boot_res_npp_evi <- boot(data = npp_evi, statistic = boot_func_npp_evi, R = 1000) # R is the number of resamples
# Compute the 95% confidence intervals for each point
ci_lower <- apply(boot_res_npp_evi$t, 2, quantile, probs = 0.025) # Lower 2.5%
ci_upper <- apply(boot_res_npp_evi$t, 2, quantile, probs = 0.975) # Upper 97.5%
# Predicted values from the original fit
evi_seq <- seq(min(abi_evi$evi_landsat), max(abi_evi$evi_landsat), length.out = 100)
pred_npp_evi <- apply(boot_res_npp_evi$t, 2, mean) # Mean of the bootstrapped predictions
# Create a data frame with predicted values and confidence intervals
pred_df_npp_evi <- data.frame(
evi_landsat = evi_seq,
npp = pred_npp_evi,
ci_lower = ci_lower,
ci_upper = ci_upper
)
lnpp_evi <- npp_evi |>
group_by(sp_code) |>
group_modify(~ {
# Ajusta el modelo
mod <- lm(npp ~ evi_landsat, data = .x)
# Tidy con coeficientes
tidy_df <- broom::tidy(mod)
# Añade R2 a cada fila
tidy_df$r.squared <- broom::glance(mod)$r.squared
tidy_df
}) |>
mutate(p = case_when(
p.value < 0.001 ~ "<0.001",
p.value >= 0.001 ~ as.character(round(p.value, 3))
)) |>
mutate(sig = case_when(
p.value < 0.001 ~ "sig",
p.value >= 0.001 ~ "no sig"
))
y_max_npp <- 1050
lnpp_evi_eq <- lnpp_evi |>
left_join(
lnpp_evi |> filter(term == "(Intercept)") |>
dplyr::select(sp_code, intercept = estimate)
) |>
mutate(
eq_text = if_else(
sig == "sig",
glue("y = {round(estimate, 2)}x {ifelse(intercept < 0, '-', '+')} {round(abs(intercept), 2)}; R² = {round(r.squared, 2)}"),
NA
)
) |>
filter(term != "(Intercept)") |>
arrange(desc(sig)) |> # ordenar por eq
rowid_to_column("id") |>
mutate(offset_y = id * 0.05 * y_max_npp,
y_max = offset_y) |>
mutate(Specie = paste0("P. ", sp_code))
npp_evi <-
npp_evi |> inner_join(
lnpp_evi_eq |> dplyr::select(sp_code, p, sig))
## Plots
# general_parameters
alpha_points <- 0.5
main_line_width <- 1.3
main_line_color <- "black"
partial_lines_width <- .85
size_points <- 2
custom_options <- list(
scale_shape_manual(
values = shape_elev,
name = "Elevation"),
scale_colour_manual(
values = colours_Specie,
labels = expression(italic("P. halepensis"), italic("P. pinaster"), italic("P. nigra"), italic("P. sylvestris")),
name = "Species"),
scale_fill_manual(
values = colours_Specie,
labels = expression(italic("P. halepensis"), italic("P. pinaster"), italic("P. nigra"), italic("P. sylvestris")),
name = "Species"),
theme_bw(),
theme(
panel.grid = element_blank(),
axis.title = element_text(size = 14, face = "bold"),
axis.text = element_text(size = 13),
axis.ticks.length = unit(.2, "cm"),
legend.title=element_text(size=14, face = "bold"),
legend.text=element_text(size=14)
)
)
plot_abi_npp <- abi_npp |>
ggplot(aes(y = abi, x = npp)) +
geom_point(aes(shape = elev_code, fill = Specie, colour = Specie), alpha = alpha_points, size = size_points) +
geom_line(data = pred_df_abi_npp, aes(x = npp, y = abi), color = "black", linewidth = 1.5) + # Fitted model
# geom_line(data = pred_df_abi_npp, aes(x = npp, y = ci_lower), color = "black", linetype = "dashed") +
# geom_line(data = pred_df_abi_npp, aes(x = npp, y = ci_upper), color = "black", linetype = "dashed") +
geom_ribbon(data = pred_df_abi_npp, aes(x = npp, ymin = ci_lower, ymax = ci_upper), alpha = 0.3) + # CI
scale_y_continuous(limits = c(0, 450)) +
ylab(expression(ABI ~ (g ~ C ~ m^2 ~ year^{-1}))) +
xlab(expression(NPP[MODIS] ~ (g ~ C ~ m^2 ~ year^{-1}))) +
custom_options
plot_abi_evi <- abi_evi |>
ggplot(aes(y = abi, x = evi_landsat)) +
geom_point(aes(shape = elev_code, fill = Specie, colour = Specie), alpha = alpha_points, size = size_points) +
geom_line(data = pred_df_abi_evi, aes(x = evi_landsat, y = abi), color = "black", linewidth = 1.5) + # Fitted model
# geom_line(data = pred_df_abi_evi, aes(x = evi_landsat, y = ci_lower), color = "black", linetype = "dashed") +
# geom_line(data = pred_df_abi_evi, aes(x = evi_landsat, y = ci_upper), color = "black", linetype = "dashed") +
geom_ribbon(data = pred_df_abi_evi, aes(x = evi_landsat, ymin = ci_lower, ymax = ci_upper), alpha = 0.3) + # CI
ylab(expression(ABI ~ (g ~ C ~ m^2 ~ year^{-1}))) +
xlab(expression(EVI[Landsat])) +
custom_options
plot_npp_evi <- npp_evi |>
ggplot(aes(y = npp, x = evi_landsat)) +
geom_point(aes(shape = elev_code, fill = Specie, colour = Specie), alpha = alpha_points, size = size_points) +
geom_line(data = pred_df_npp_evi, aes(x = evi_landsat, y = npp), color = "black", linewidth = 1.5) + # Fitted model
# geom_line(data = pred_df_npp_evi, aes(x = evi_landsat, y = ci_lower), color = "black", linetype = "dashed") +
# geom_line(data = pred_df_npp_evi, aes(x = evi_landsat, y = ci_upper), color = "black", linetype = "dashed") +
geom_ribbon(data = pred_df_npp_evi, aes(x = evi_landsat, ymin = ci_lower, ymax = ci_upper), alpha = 0.3) + # CI
ylab(expression(NPP[MODIS] ~ (g ~ C ~ m^2 ~ year^{-1}))) +
xlab(expression(EVI[Landsat])) +
custom_options
### By species
p_sp <- (
(plot_abi_npp +
geom_smooth(aes(linetype = sig, group = Specie, colour = Specie),
linewidth = partial_lines_width, method = "lm", se = FALSE) +
scale_linetype_manual(values = lines_lm, guide = "none") +
scale_x_continuous(
limits = c(150,1050),
breaks = c(150,300,450,600,750,900,1050)) +
annotate(geom = "text", label = "a", x = 150, y = Inf, hjust = .2, vjust = 1.5, size = 7, fontface = "bold") +
geom_text(data = labi_npp_eq |> filter(!is.na(eq_text)),
aes(x = 600, y = y_max, label = eq_text, colour = Specie),
hjust = 1.1, vjust = 1.1, size = 2.75,
fontface = "bold", show.legend = FALSE)) +
(plot_abi_evi +
geom_smooth(aes(linetype = sig, group = Specie, colour = Specie),
linewidth = partial_lines_width, method = "lm", se = FALSE) +
scale_linetype_manual(values = lines_lm, guide = "none")+
scale_x_continuous(
limits = c(0.1,0.5),
breaks = c(0.1, 0.2, 0.3, 0.4, 0.5)) +
annotate(geom = "text", label = "b", x = .1, y = Inf, hjust = .2, vjust = 1.5, size = 7, fontface = "bold") +
geom_text(data = labi_evi_eq |> filter(!is.na(eq_text)),
aes(x = Inf, y = y_max, label = eq_text, colour = Specie),
hjust = 1.1, vjust = 1.1, size = 2.75,
fontface = "bold", show.legend = FALSE)) +
(plot_npp_evi +
geom_smooth(aes(linetype = sig, group = Specie, colour = Specie),
linewidth = partial_lines_width, method = "lm", se = FALSE) +
scale_linetype_manual(values = lines_lm, guide = "none") +
scale_x_continuous(
limits = c(0.1,0.5),
breaks = c(0.1, 0.2, 0.3, 0.4, 0.5)) +
scale_y_continuous(
limits = c(0,1050),
breaks = c(0,150,300,450,600,750,900,1050)) +
annotate(geom = "text", label = "c", x = .1, y = Inf, hjust = .2, vjust = 1.5, size = 7, fontface = "bold") +
geom_text(data = lnpp_evi_eq |> filter(!is.na(eq_text)),
aes(x = 0.5, y = y_max, label = eq_text, colour = Specie),
hjust = 1.1, vjust = 1.1, size = 2.75,
fontface = "bold", show.legend = FALSE))
) & theme(legend.position = "bottom", legend.text = element_text(size = 14))
compare_plot_sp <- p_sp + plot_layout(guides = "collect") & guides(
shape = guide_legend(override.aes = list(stroke = .5, size = 3, colour = "black")),
fill = guide_legend(override.aes = list(size = 3))) & theme(legend.box = "vertical")
compare_plot_sp
```
```{r}
#| echo: false
ggsave(
compare_plot_sp,
file = here::here("output/plot_compare_abi_npp_evi_sp.png"),
dpi = 400,
width = 7.09*1.9, height = 7.09*.8
)
# ggsave(
# compare_plot_sp,
# file = "output/compare_abi_npp_evi_sp.pdf",
# dpi = 400,
# width = 7.09*2.2, height = 7.09*1.8/2
# )
ggsave(
compare_plot_sp,
file = here::here("output/plot_compare_abi_npp_evi_sp.svg"),
dpi = 400,
width = 7.09*1.9, height = 7.09*.8
)
```